Tutorial - How to install and load the Identifier Mapping Service with data needed for gene-to-variant and variant-to-gene
This tutorial explains how to install and run a BridgeDb Identifier Mapping Service (IMS) with gene-to-variant and variant-to-gene functionality.
Step 1. Downloading the gene-variant mappings
Download the linksets data you want to load into the IMS from http://bridgedb.org/data/gene_database/linkset/.
Step 2. Install Docker
A prerequisite is a Docker installation. You can download Docker from various places and many GNU/Linux distributions ship a version as part of their distribution. Other options include:
- Docker Community Edition: https://www.docker.com/get-docker
- Docker Toolbox for older Mac and Windows versions: https://docs.docker.com/toolbox/overview/
- Docker for Windows: https://docs.docker.com/docker-for-windows/
Step 3. Downloading the BridgeDb IMS Docker
Pull the IMS docker image:
docker pull openphacts/identitymappingservice
Step 4. Set up the MySQL service
Create the mysql docker image to store the linksets:
docker run --name mysql-for-ims -e MYSQL_ALLOW_EMPTY_PASSWORD=yes \
-e MYSQL_DATABASE=ims -e MYSQL_USER=ims -e MYSQL_PASSWORD=ims \
-d mysql
Step 5. Loading the gene-gene and gene-variant link sets
The next command line example assumes that the file load.xml is present the /home/johndoe/data5 directory.
Unix:
docker run --link mysql-for-ims:mysql -v /home/johndoe/data5:/staging \
openphacts/identitymappingservice loader file:///staging/load.xml
MINGW64 / Windows:
docker run --link mysql-for-ims:mysql -v /c/Users/johndoe/data5:/staging openphacts/identitymappingservice loader file:///staging/load.xml
An example of the load.xml file (more info):
<?xml version="1.0"?>
<loadSteps>
<clearAll/>
<void>file://staging/Ensembl_SNP_dataset.void.ttl</void>
<linkset>file:///staging/PolyPhen_91.ttl</linkset>
</loadSteps>
Step 6. Run the IMS docker image
docker run --name ims --link mysql-for-ims:mysql -p 8081:8080 -d openphacts/identitymappingservice
You can confirm the instance is running with:
docker ps
Both these commands will give you a long identifier which you should remember for later.
Step 7. Check the QueryExpander expose at the local port 8081
Open the following webpage in a browser:
It should look like this:
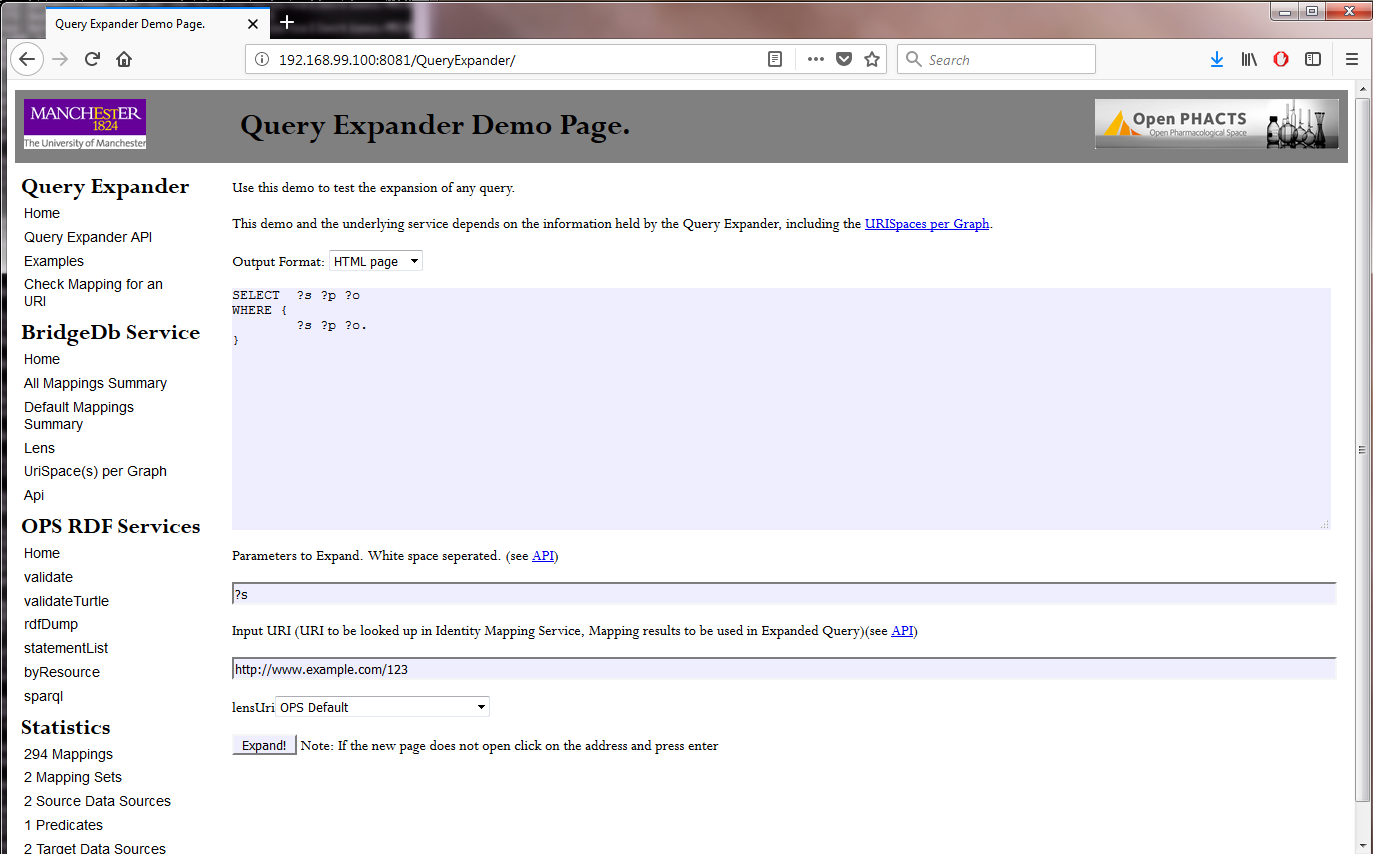
Step 8. Shutting down the Docker image
Shutting down the docker is done with the command docker stop and the identifier you got with the docker run or docker ps commands, for example:
docker stop b5e9715818f6
of
docker stop ims
Where b5e9715818f6 or ims happens to be the ID/name for the running image for the author at the time of writing.
You can delete the images with:
docker container rm b5e9715818f6 d989788c0b03
Where the two IDs are the one for the MySQL-IMS docker and the IMS docker containers.